Investigating the causes and distribution of bacterial gastroenteritis within the UK population
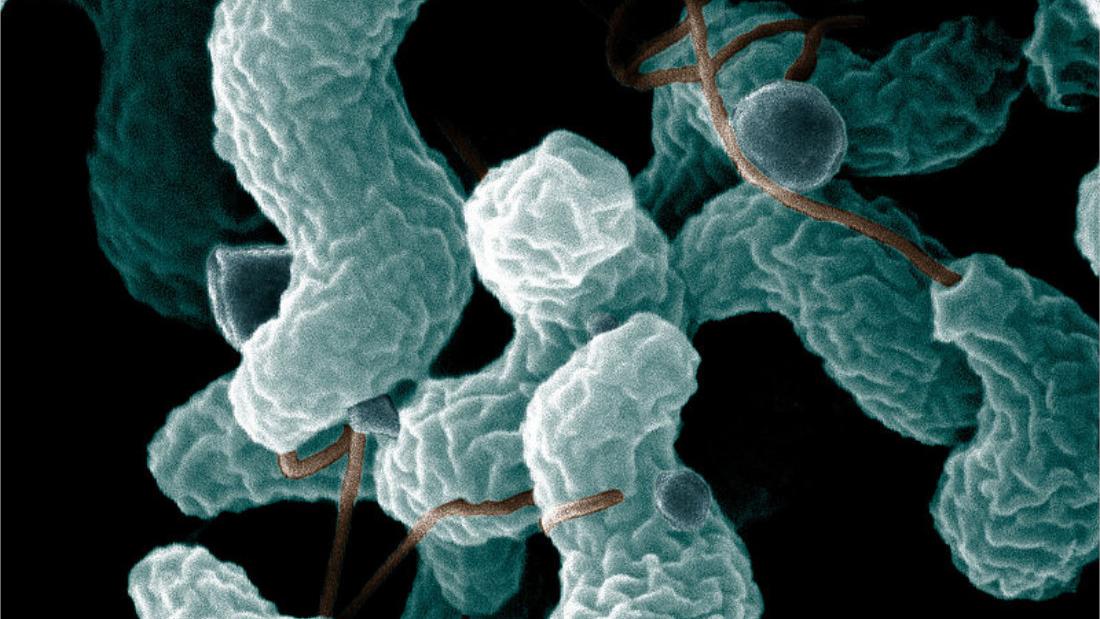
Campylobacter bacteria cause mild to severe food poisoning or gastroenteritis, amongst other symptoms. The cost of this is large and has been estimated to be as much as £50 million anually in the UK alone. Controlling Campylobacter in the food chain is critical to reducing the incidence of this disease. Our data looks at the two species of Campylobacter bacteria which make humans sick most often.
There are 35 species of Campylobacter but only two of these make humans sick regularly. In the UK, 90% of cases are caused by Campylobacter jejuni with the other 10% caused by Campylobacter coli. Both of these species are found found naturally in domestic and wild animals. From there they can infect us through the food we eat mainly meat or through contaminated water or milk.
Campylobacter lives naturally and apparently harmlessly in the guts of birds and mammals. In scientific terms we say that they are members of the commensal gastrointestinal microbiota of these animals.
Infections are passed to humans either from direct contact with contaminated faeces or through indirect transmission via contaminated food or drink.
By studying the incidence of sickness, investigating outbreaks, and experimentation the scientific community has identified contaminated chicken meat as a major cause. You can use the same data that we use to look at the causes of infection in people.
These data have been collected as part of a long-term study of Campylobacter infections in Oxfordshire, funded in MaidenLab since 2003 by the Department for Environment Food and Rural Affairs (DEFRA) and the Food Standards Agency (FSA).
In the UK, the incidence of Campylobacter infection varies with age, being highest among those under five years of age. Males are more frequently affected, with a male-to-female ratio of 1.2 to 1.
The diagram below shows the relationship between gender and Sequence Type Clonal Complexes.
Bacteria can be hard to categorise within a species as they have very large populations, are very diverse, and can evolve quickly. Using genomic sequence data we can group sub-species together to help us better understand their distribution and evolution. We use about 450-500 internal stable sequences (found around housekeeping genes) and then group broadly similar results together into sequence types. These sequence types (STs) are then grouped into families which we call clonal complexes (ccs).
The numbering of clonal complexes comes from the predominant sequence type found in the family. For example, the sequence type (ST-21) is most predominant in its grouping and hence the clonal complex is also called cc-21.
Text about findings in species
Clonal complexes 21, 48 and 353 are the most frequently occured Campylobacter jejuni, while clonal complex 828 is the most frequently occuring Campylobacter coli. Click the ribbons within the visualisation below to see some recent patient data.
Reference: Campylobacter genotypes from food animals, environmental sources and clinical disease in Scotland 2005/6 Sheppard, S. K., Dallas, J. F., MacRae, M., McCarthy, N. D., Sproston, E. L. , Gormley, F. J., Strachan, J. C., Ogden, I. D., Maiden, M. C. J. and Forbes, K. J., International Journal of Food Microbiology, 134 (2009), pp 96-103.
Publications: You can find the full list of related publications on PubMed
We would like to acknowledge support from:-
- Food Standards Agency funds the Campylobacter Surveillance Project.
- Public Health England collaborates in the generation of these data.
- The Wellcome Trust funds the PubMLST.org website which hosts these data.
- National Institute of Health protection Unit in Gastrointestinal Infections (NHIR HPRU-GI)
We gratefully acknowledge Valérie Lavigne, Marius Panga and Jayaram Shivas Vadakumpuram for their kind permission to allow us to use and build upon their work.





